 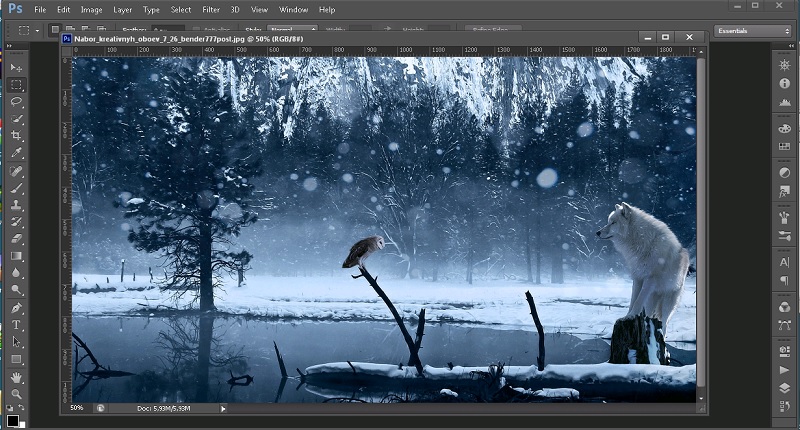
The structure of the unactivated form of ribulose-1,5-bisphosphate carboxylase/oxygenase was refined at a resolution of 2.0 A to an R-factor of 17.1%. While it would seem probable that these similarities represent divergent evolution from a primordial heme-binding protein, the possibility of structural convergence to a functionally satisfactory protein cannot be excluded. The roughly 50 residues missing at the carboxy end of the known cytochrome b5 fragment could correspond in part to the H helix in the globins. The beta-sheet of cytochrome b5 is inserted into a corresponding cavity of the globins forming an additional lining to the heme pocket. The heme itself is rotated by 53 degrees about its normal but such a change is energetically minimal and conservative as the heme side groups are not directly involved in the function of the molecules. Larger differences in structure are observed on the distal side of the heme, coincident with the most changeable part of the globin structures. The proximal histidine of the globins and two adjacent helices are equivalent to the sixth iron ligand and adjacent helices of cytochrome b5. When these proteins have been superimposed, the heme irons are found to be less than 1.4 A separated and the heme normals are inclined by less than 9.5 degrees. Of the 85 three-dimensionally characterized residues of cytochrome b5, 51 are found to be structurally and topologically equivalent to the globin fold. Energy computation suggests that this mutation might increase the carboxylation activity. The mutant Leu335→Arg of rice Rubisco has been designed using the graphics software TURBO-FRODO and the program package XPLOR. The activesite structures are basically the same for the two species. The main-chain structure of rice Rubisco does not show obvious difference from that of spinach Rubisco, and the side-chains are located in the most favorable positions and orientations. Mg2+♼ABP of rice Rubisco has also been modeled and optimized using the program package XPLOR.The three-dimensional structure of the quaternary complex Rubisco 
The molecular modeling was carried out in the following procedure: amino-acid sequence substitution manual adjustment of the initial model and energy minimization. Molecular model of rice Rubisco has been constructed by molecular modeling method on a computer graphics workstation using graphics software TOM-FRODO and QUANTA/CHARMm, based on the three-dimensional structure of spinach Rubisco. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
